Technologies of CLST
ACCIS: Automated Cell Culturing System
Co-developed with the Hamilton Robotics, we designed, built and integrated Automated Cell Culturing System (ACCIS). ACCIS has a robotic arm, a centerpiece that integrates multiple devises into one system by transporting culture dishes to each component. The system also includes two large CO2 incubators that holds up to 500x 96-well plates, one carousel for plate loading, Celigo S imaging system, BioTek 406 plate washer, and a MicroStar liquid handler (Figure 1).
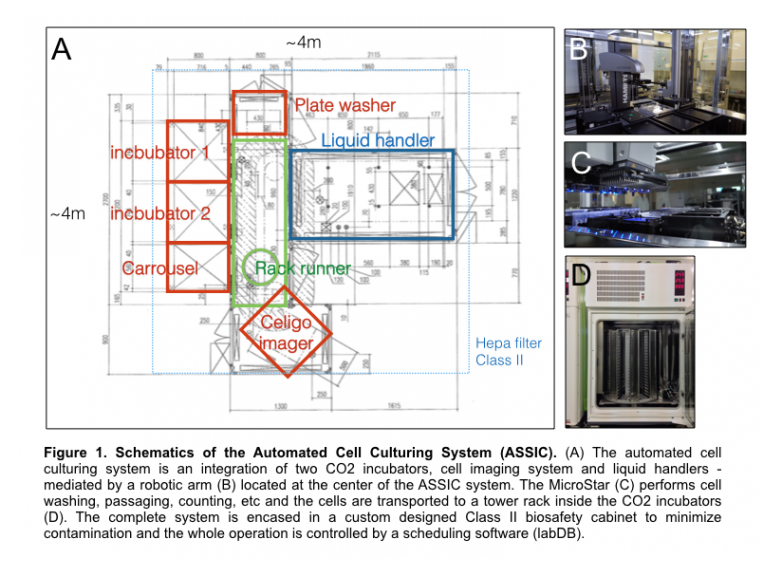
ACCIS is uniquely encased in a custom designed Class II Biosafety cabinet, which allows for the production and transduction of lentivirus and provides extra safety and cleanliness for most of mammalian cell culture activities.
The heart of ACCIS is the software that communicates with all devices and commands instructions to the rest of the system. LabDB, designed to run ACCIS, is a scheduler to instruct when-and-how to initiate, and it also has the capability to “decide” whether to continue or return based on the cell growth rate.
The automation naturally allows for large-scale cell culturing. ACCIS can culture and propagate hundreds of cultured cells without a need of human intervention. In addition, ACCIS is designed to transfect/transduce genetic materials to manipulate gene expression inside the cell. This technique is essential to decipher the function of a gene and we are currently using ACCIS to knockdown thousands of long non-coding RNAs (lncRNA) in various human cell types, including human induced pluripotent stem (hIPS) cells.
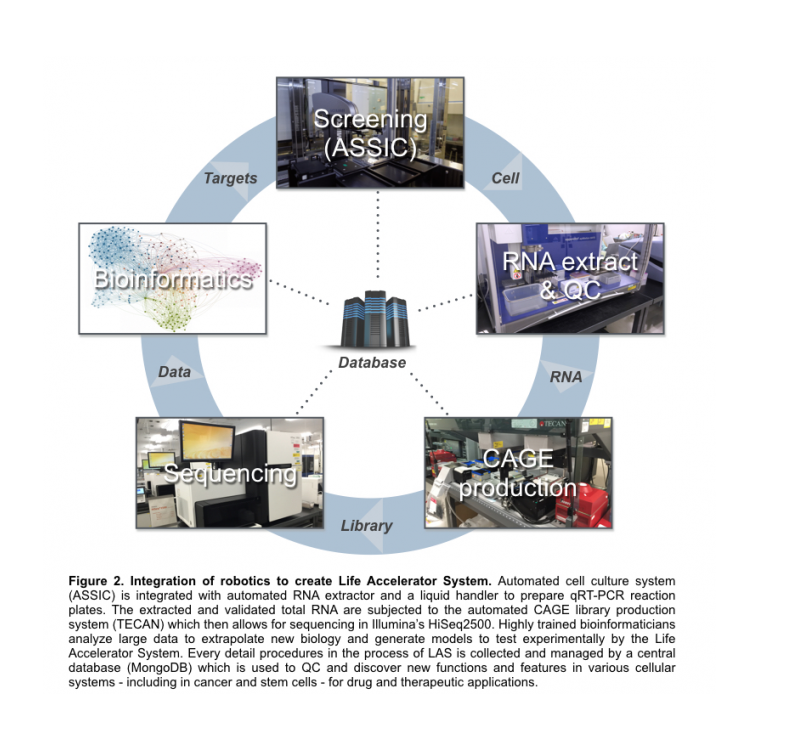
In DGT, we are continuously making strategic efforts to expand-and-integrate robotics to create a systematic pipeline to functionally annotate genes. Hence, the Life Accelerator System (LAS) was designed as a master pipeline for FANTOM6. The LAS integrates ASSIC in conjunction with two other epMotion’s Liquid Handling systems for RNA extraction and expression validation, TECAN Robotics for high-throughput CAGE library production, and Illumina’s HiSeq2500 Next Generation Sequencer for molecular phenotyping. Each component of LAS is connected to a central database that collects, organizes, and disseminates meta-data for quality control and data analysis performed by specialized bioinformaticians in DGT (Figure 2).